
Figure 1. All of Us Research Program Research Priorities Workshop (March 2018).
Unless you have been living under a rock, you know that NIH has launched the largest effort to date to establish a cohort in the United States. Chatter about the need for a nationally representative cohort bubbled up more than ten years ago in a commentary by Dr. Francis Collins (PMID:15164074), then head of the National Human Genome Research Institute. After a bit of back and forth in the literature in support (PMID:17230172) and against (PMID:17230171) such an effort, not much happened. Meanwhile, the smaller, more specialized cohorts quietly embraced the motto “keep being you” and soldiered on alone and in collaborations as the golden age of genome-wide association studies (GWAS) unfolded.
Flash forward more than a decade, and much has changed. A change in NIH leadership, a new administration, and sprinkle in ARRA (with the HITECH Act) and a little UK BioBank envy and voila, the US cohort is reborn!
The rebirth of the US cohort is in the form of the Precision Medicine Initiative Cohort Program, rebranded as All of Us (Figure 1). This is not your mom’s cohort. Technically, it is designed to represent but not necessarily be population-based. And, the data collection aims to be thoroughly modern: electronic health records, syncforscience, and other data portal opportunities. There’s an app for that, indeed!

Figure 2. All of Us Research Program Research Priorities Workshop (March 2018).
So much of what All of Us proposes to do has never been done before at such a large scale or at the pace. Although All of Us has been called out for being slower than originally planned (PMID:28883052), it’s still organizing at a fast pace to the point that they are riding the train while still laying the tracks. To ensure the train doesn’t derail soon after leaving the station, the All of Us program organized a Research Priorities Workshop in March of 2018 (Figure 2). As the title of the workshop implied, the major goals of the meeting centered on research priorities and the data required to answer specific scientific questions of interest. Investigators invited to or just interested in All of Us were encouraged prior to the workshop to submit online “use case” scenarios designed to capture research questions, data needed, data collection methods, and data collection specifications.

Figure 3. Bethesda in March–it’s like I’m in Cleveland (2018)!
Although our institutions here in Cleveland are not directly or officially involved in All of Us, I was fortunate enough to be invited to the in-person workshop. The three-day in-person meeting was organized by health conditions and cross-cutting themes, and each workshop participant was asked to select one area for each. As a generalist, I struggled with health conditions, so I selected “cardio-respiratory and blood” given my past work on lipid genetics. For my cross-cutting theme, I chose “Informatic, Methodologies, Ethical/Legal, and Statistical Research” over the perhaps more obvious “Genomics and other Omics.” Although my selected theme was a hodgepodge of themes not assigned to another group, I anticipated that this group would offer interesting insights on the directions All of Us might be taking in biomedical informatics given an expected component will be electronic and personal health records.
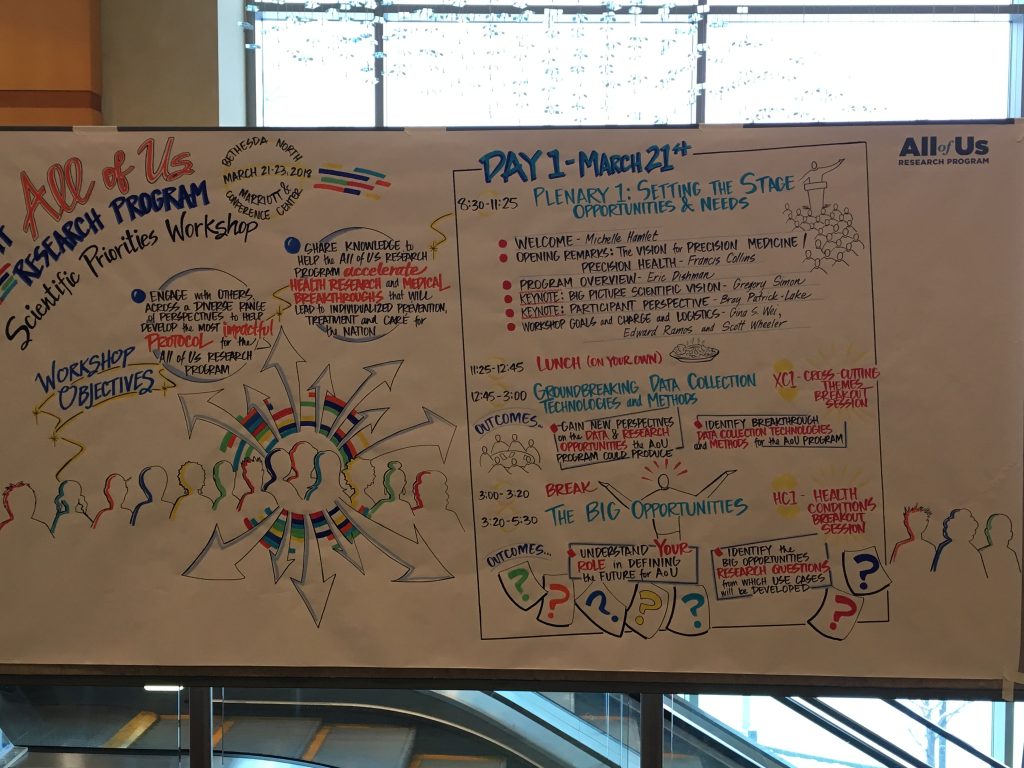
Figure 4. Day 1 of the All of Us Research Program Research Priorities Workshop (March 2018).
Despite the weather (Figure 3) and travel chaos that ensued, the workshop was highly organized and each group was led by obviously experienced moderators. Our moderator was Megan (Meg) Doerr, Principal Scientist at SAGE Bionetworks in Seattle, WA. It turns out that prior to SAGE, Meg was a genetic specialist at the Cleveland Clinic. Small world! She was also a teacher at a K-12 school in her previous life, which explains her effective moderating skills.
Starting with Day 1 (Figure 4), each group session was further split into small groups of four or five investigators sitting at round tables with assigned tasks. I sat at table 3’s “Elucidating Disease Mechanisms” with Drs. Svati Shah (Duke University), Soumitra Sengupta (Columbia University), and Jeff Whittle (Medical College of Wisconsin). We were later joined by Olayiwola Ayodeji, a research analyst with All of Us (Figure 5). Both Drs. Shah and Wittle are physicians, so they had a leg up on specific research questions related to disease mechanisms. To boot, Dr. Shah is PI of Project Baseline, a smaller version of All of Us of sorts organized and funded by Duke University School of Medicine, Stanford Medicine, Verily, and Google. Dr. Shah stressed that the basic protocols for All of Us had already been written, so useful use cases would be those that could lead to protocol revisions or new protocols.
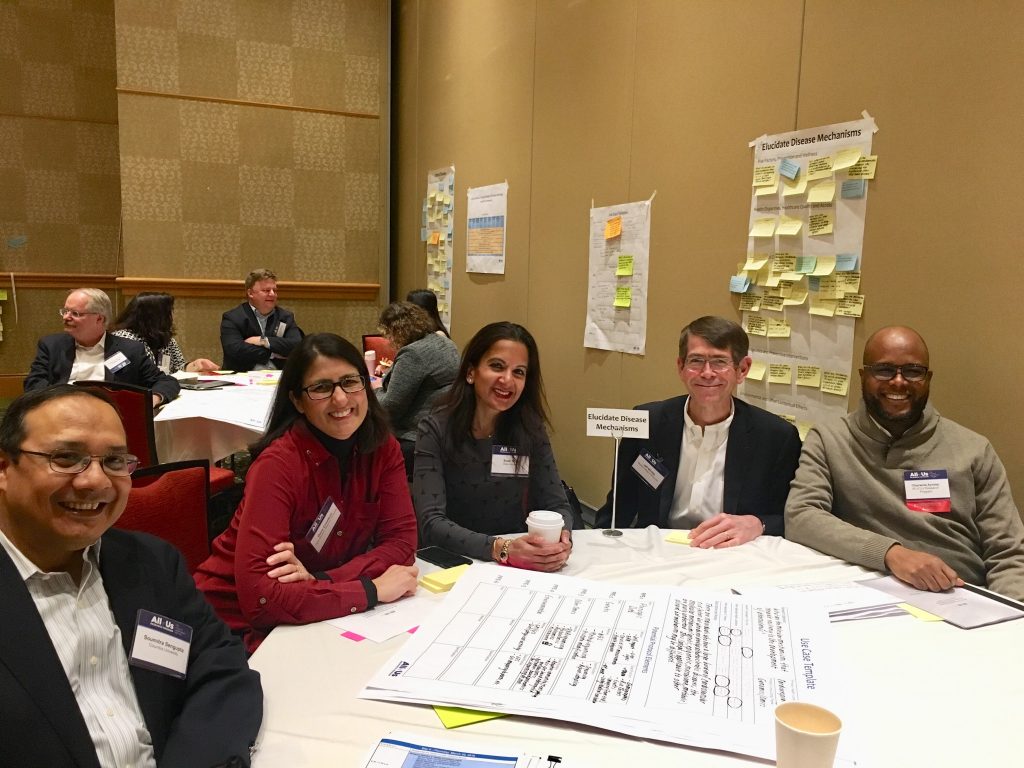

Figure 5. Table 3’s Drs. Soumitra Sengupta, Dana Crawford, Svati Shah, Jeff Whittle along with All of Us analyst Olayiwola (Wally) Ayodeji at the All of Us Research Program Research Priorities Workshop (March 2018).
While Dr. Sengupta and I did not have much to add to specific research questions related to disease mechanisms, we did endorse the need for data harmonization and complete longitudinal data for electronic health records. Pretty much everyone was in agreement with those issues. Of course, genomic discovery studies are a favorite of mine, but Dr. Shah correctly pointed out that those will certainly be done and can be done with existing protocols. Focus, Crawford, focus!

Figure 6. Dr. Svati Shah presenting the table’s use cases at the All of Us Research Program Research Priorities Workshop (March 2018).
The workshop culminated with each small group presenting their use cases. Dr. Svati Shah represented our table in a tour de force of research ideas (Figure 6). As if that weren’t impressive enough, we (“we” being Dr. Shah) won best handwriting (Figure 7)!

Figure 7. Our group received an A for best handwriting at the All of Us Research Program Research Priorities Workshop. Thanks, Dr. Svati Shah (March 2018)!
All use cases from all groups across the workshop were collected and entered into the All of Us database (Figure 8). We’re not quite sure what became of them beyond the meeting, but there will certainly be analyses behind the scenes to identify new research priorities and needed data collection or analysis methodologies.
Soon after the workshop, NIH announced the official launch of All of Us for May 6, 2018. And, as of the first newsletter in September 2018, All of Us has registered approximately 100,000 participants. Regardless of where you stand on the usefulness of the proposed cohort or its feasibility, the train has left the station. Let’s see where it takes us.

Figure 8. Days 2 and 3 of the All of Us Research Program Research Priorities Workshop (March 2018).
